 
Nucleosomes are spaced at intervals of about 200 nucleotides pairs along the DNA strand, which may explain why new Okazaki fragments are synthesized on the lagging strand at intervals of 100-200 nucleotides in eukaryotes, instead of 1000-2000 nucleotides as in bacteria.Īs already discussed by me and in the comments, increasing telomere longevity in eukaryotes by reducing the length of okazaki fragments, and consequently reducing the part of telomere cut-off after every division might be another reason for having shorter fragments in eukaryotes as compared to prokaryotes which do not contain telomeres.ĮDIT : Having thought over it again, the relation between Okazaki fragment length and telomeres do not seem logical as only RNA primer length is important when considering telomere length.īalakrishnan and Bambara (2013) explain the regulation of (and differences between) prokaryotic and eukaryotic Okazaki fragments in detail. The scientists found there was a discontinuous replication process by pulse-labeling DNA and observing changes that pointed to non-contiguous replication.I have found one more possible reason from Bruce Alberts' The Molecular Biology of the Cell: (Ch. They are: substrates, template, primer and enzymes. However, the size and number of the Okazaki fragments can be manipulated by varying the activity of gp61, the gp45 and gp44/62 levels, and the rate of synthesis by the lagging-strand gp43 to create a pattern of gapped Okazaki fragments, i.e. Before this time, it was commonly thought that replication was a continuous process for both strands, but the discoveries involving E. 640K views 3 years ago Biology This biology video tutorial provides a basic introduction into DNA replication. Requirements for DNA Synthesis There are four basic components required to initiate and propagate DNA synthesis. During the 1960s, Reiji and Tsuneko Okazaki conducted experiments involving DNA replication in the bacterium Escherichia coli. Despite attachment to a sliding clamp, the polymerase on the lagging strand must cycle on and off DNA for each Okazaki fragment. The entire replication process is considered "semi-discontinuous" since one of the new strands is formed continuously and the other is not. Once the fragments are made, DNA ligase connects them into a single, continuous strand. The primase and polymerase move in the opposite direction of the fork, so the enzymes must repeatedly stop and start again while the DNA helicase breaks the strands apart. Okazaki fragments are around 100-1000 nucleotides long whereas lagging strand can be more than 10,000 nucleotides long. This causes periodic breaks in the process of creating the lagging strand. The main difference between Okazaki fragments and lagging strand is that Okazaki fragments are much shorter than lagging strand. 
The synthesis of a new DNA molecule may lead to some errors, which, if not corrected, would lead to mutations in the genome. The lagging strand, however, cannot be created in a continuous fashion because its template strand has 5’ to 3’ directionality, which means the polymerase must work backwards from the replication fork. Lagging strand synthesis requires sequential action of series of enzymes that initiate, elongate, and join pieces of DNA of about 1000 nucleotides called Okazaki fragments. Strand-displacement synthesis occurs, whereby. 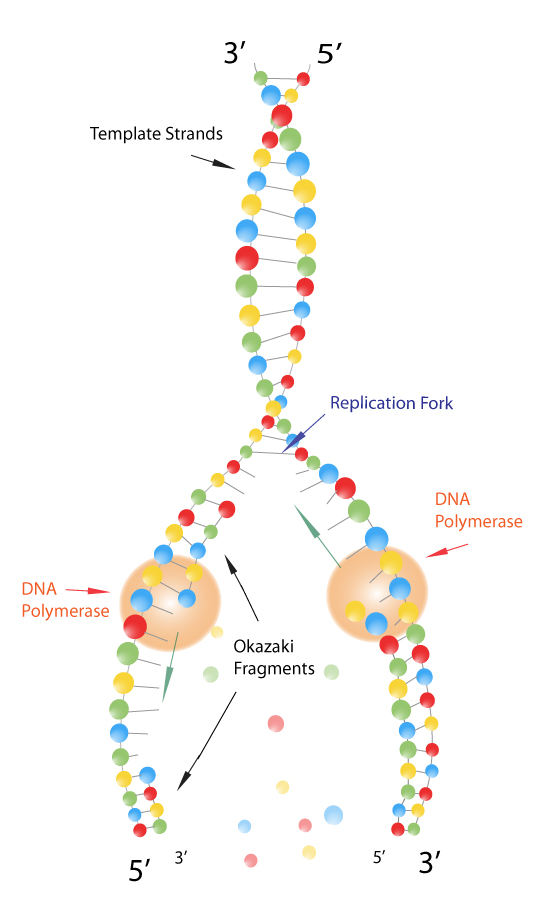
One strand, the leading strand, undergoes a continuous replication process since its template strand has 3’ to 5’ directionality, allowing the polymerase assembling the leading strand to follow the replication fork without interruption. Short DNA fragments, about 100 bases long, called Okazaki fragments are synthesized on the RNA-DNA primers first. Because these enzymes can only work in the 5’ to 3’ direction, the two unwound template strands are replicated in different ways. Following this fork, DNA primase and DNA polymerase begin to act in order to create a new complementary strand. Transient components of lagging strand of DNA Asymmetry in the synthesis of leading and lagging strandsĭuring DNA replication, the double helix is unwound and the complementary strands are separated by the enzyme DNA helicase, creating what is known as the DNA replication fork. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
